Temporal Data Requirement for Predicting Unplanned Hospital Readmissions
Source: arXiv:2605.00738 · Published 2026-05-01 · By Ramin Mohammadi, Vahab vahdat, Sarthak Jain, Amir T. Namin, Ramya Palacholla, Sagar Kamarthi
TL;DR
This study addresses the critical question of how much historical Electronic Health Record (EHR) data is needed to optimally predict 30-day unplanned hospital readmissions following hip and knee arthroplasty surgeries. While prior work assumed that using more historical data generally improves machine learning prediction, this research systematically analyzes time windows ranging from the day of surgery up to three years prior. The dataset includes over 4 million structured encounter records and 80,000 clinical notes from 7,174 patients. The authors extract features from structured data as well as unstructured clinical notes using multiple encoding methods, both traditional (Bag of Words, TF-IDF, LDA) and neural (BERT, 1D CNN, BiLSTM). They then train and evaluate predictive models on clinical notes only, structured data only, and combinations of both.
The key novelty is the discovery that the optimal temporal window varies by modality: clinical notes achieve maximum predictive power using only 3 to 6 months of prior data, whereas structured data benefits from longer windows but plateaus after 12 months. This holds consistently across all encoder architectures and modeling variants including multitask learning. Combining both modalities yields performance trends similar to clinical notes alone, indicating the high information value in recent free text. These findings challenge the simplistic "more data is always better" assumption for temporal EHR data in readmission prediction, and provide concrete guidelines on data retention to optimize model accuracy while minimizing processing burden.
Key findings
- Optimal clinical notes observation window for 30-day readmission prediction is 3 to 6 months prior to surgery.
- Structured data model performance increases with observation window length but plateaus after 12 months.
- Combining clinical notes and structured data yields performance trends similar to clinical notes alone, emphasizing recent unstructured text's predictive power.
- Trends in temporal data requirements hold across multiple encoding methods including BERT, 1D CNN, BiLSTM, BoW variants, TF-IDF, LDA.
- Multitask learning models jointly predicting hip and knee readmissions show improved performance but same temporal data pattern as independent models.
- Using only surgical notes from the day of surgery underperforms models that also incorporate the last 3 months of historical data for both clinical notes and structured data.
- Patient cohort: 7,174 patients with 4M+ encounter records and 80K clinical notes; hip readmission positive rate ~9.5%, knee ~10.5%.
- Maximum AUROC achieved by combined clinical notes + structured data models was approximately 0.93 for knee procedures using 3-month history.
Methodology — deep read
Threat Model & Assumptions: The study assumes a predictive modeling context for 30-day hospital readmissions where the 'adversary' concept is not directly applicable. The problem is framed as a supervised learning task with labeled cases indicating presence or absence of unplanned inpatient readmission within 30 days post hip or knee arthroplasty. The models do not assume future data availability beyond the observation window specified. Data partitioning ensures patient-level splits to avoid information leakage.
Data Provenance & Characteristics: Data was retrospectively collected from the Partners Healthcare EHR system, specifically Epic, covering surgeries from 2006-2016. Initial cohort included 10,534 patients; after excluding deceased, older than 90, or without clinical notes, 7,174 patients remained. Structured data encompassed over four million encounter records across eight feature domains including demographics (10 features), labs, medications, diagnoses, procedures, vitals, and admission info (total 54 variables). Unstructured data included 80,000 clinical notes, primarily free-text narratives from physicians and nurses, also containing some tabular data parsed. Data labels were binary: positive class represented patients readmitted within 30 days post-surgery; negative otherwise. The positive class prevalence was low (around 9.5% hip, 10.5% knee), necessitating class imbalance handling via weighting, oversampling, and undersampling.
Feature Extraction and Encoding: Structured data were directly encoded as numeric or categorical variables, with categorical features represented as indicators or embeddings. Clinical notes required textual encoding: multiple non-neural methods (binary/count Bag of Words, TF-IDF, LDA topic distributions) and neural methods (BERT contextual embeddings, 1D CNN with multiple kernel sizes, BiLSTM, simple average of embeddings) were implemented. Neural encoders used 300-dimensional word embeddings (initialized with pretrained vectors), and outputs aggregated using max-pooling or attention-weighted sum. A hierarchical BiLSTM with attention over notes was also tested. The clinical BERT model was fine-tuned on this task using the Clinical BERT variant pretrained on MIMIC III.
Modeling and Training: Models were logistic regression variants (standard, L1, L2 regularized) atop non-neural text features, and neural architectures for the neural encoders. Separate predictive models were trained independently for hip and knee surgeries, and multitask models sharing encoders were explored to leverage shared structure. Training split was 70% train, 15% validation, 15% test at patient-level, preserving demographic distribution. Class imbalance was addressed using multiple strategies, though the final choice is not fully detailed. Hyperparameters included embedding dimension 300, BiLSTM hidden size 256, CNN 256 filters across kernel sizes. Models were run for multiple epochs until convergence, details on optimizer, learning rate, batch size, or random seed strategy were not explicitly specified in the paper.
Evaluation Protocol: Models were evaluated primarily by Area Under the Receiver Operating Characteristic Curve (AUROC). Confidence intervals (95%) were reported across model variants. Evaluation was performed separately on validation and test sets. Ablations included analyses across different time windows of historical data (from 0 months day of surgery to 36 months prior), across data modalities (clinical notes, structured data, combined), across model architectures (non-neural vs neural encoders), and independent vs multitask learning. Results were consistent across hip and knee cohorts. No explicit statistical tests beyond confidence intervals were mentioned. Distribution shift or external validation beyond this single-site dataset was not performed.
Reproducibility: The raw EHR dataset itself is not publicly available due to patient privacy constraints. Code release or pretrained model weights were not stated. Logical steps for feature encoding and model training are described in detail, but exact parameter settings and training procedures are partially omitted. The use of standard models like BERT and logistic regression facilitates some reproducibility given access to similar data. A concrete example of the workflow: For clinical notes with a 3-month lookback window before surgery, BERT embeddings were extracted and passed through a trained BiLSTM encoder with attention, then combined with structured data features encoded as vectors, and jointly fed to a logistic regression classifier predicting 30-day readmission label. Performance was measured as AUROC on the held-out test split, demonstrating a peak value near 0.93.
Technical innovations
- Systematic investigation of temporal observation window length for structured and unstructured EHR data in hospital readmission prediction.
- Comprehensive comparison across multiple text encoding schemes including classical BoW, LDA and advanced neural encoders (BERT, 1D CNN, BiLSTM with attention).
- Demonstration that the optimal historical data window differs by data modality: short window (3-6 months) for clinical notes vs longer window (up to 12 months plateau) for structured data.
- Application and comparison of multitask learning architectures sharing encoders to jointly model hip and knee readmission prediction while validating temporal window findings.
Datasets
- Internal Partners Healthcare EHR dataset — 7,174 patients, 4 million encounter records, 80,000 clinical notes — proprietary, not publicly available
Baselines vs proposed
- Non-neural encoder (e.g., TF-IDF logistic regression) clinical notes only: AUROC peaks around 0.88 at 3-6 months; structured data only model peaks at ~0.69 at 12 months.
- Neural encoder (e.g., BERT + BiLSTM) clinical notes only: AUROC ~0.90 at 3-6 months vs structured data only model ~0.69 at 12 months.
- Combined clinical notes + structured data with neural encoders: AUROC reaches ~0.93 for knee surgeries at 3 months window, outperforming either modality alone.
- Multitask model combining hip and knee readmission prediction achieves similar temporal window performance patterns with slight AUROC gains vs independent models.
Figures from the paper
Figures are reproduced from the source paper for academic discussion. Original copyright: the paper authors. See arXiv:2605.00738.
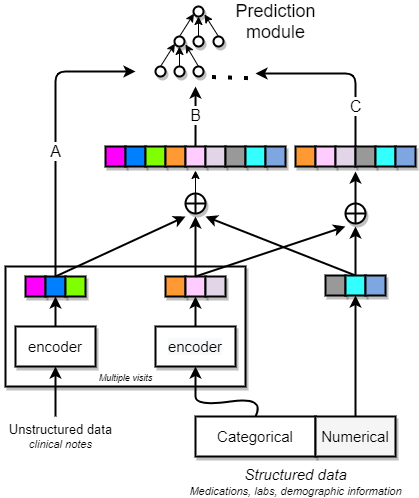
Fig 1: A schematic of feature encoding. Structured data contained both categorical and
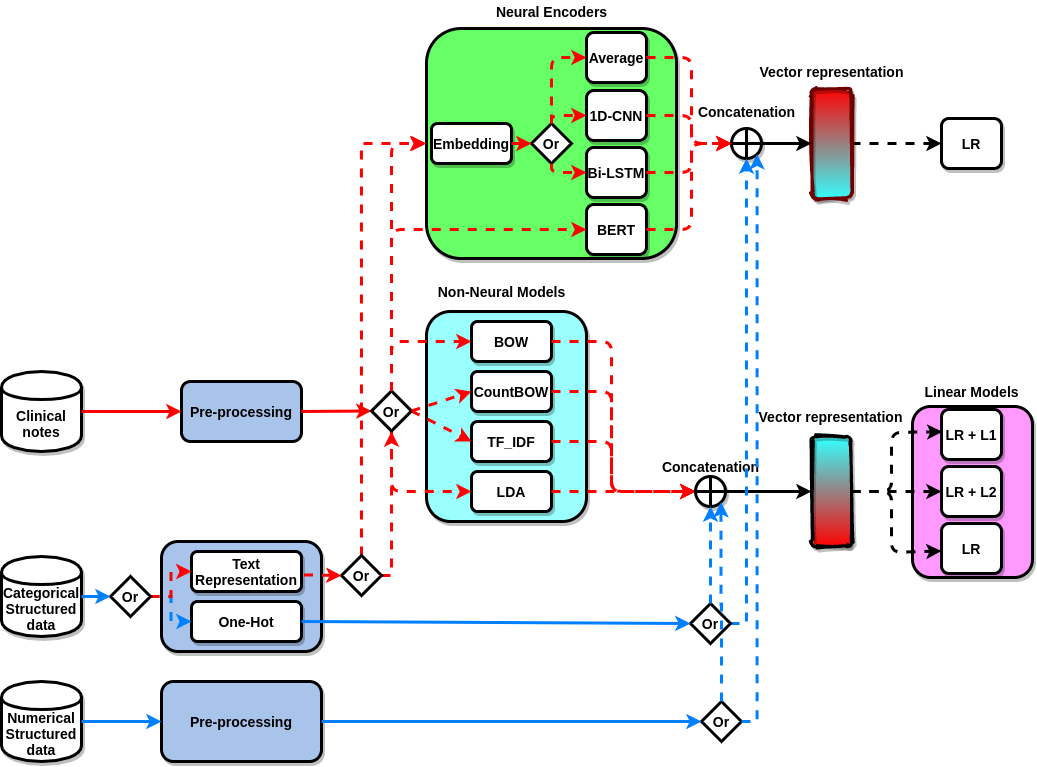
Fig 2: A schematic of model development, representing three sources of data, clinical
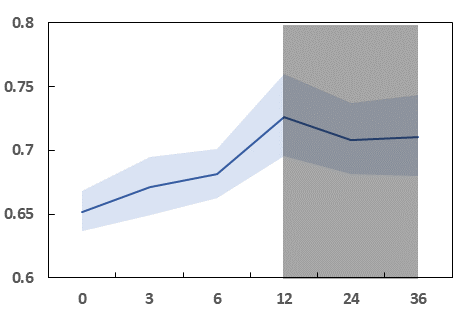
Fig 3: The variation in performance (mean and 95% confidence interval) of AUROC
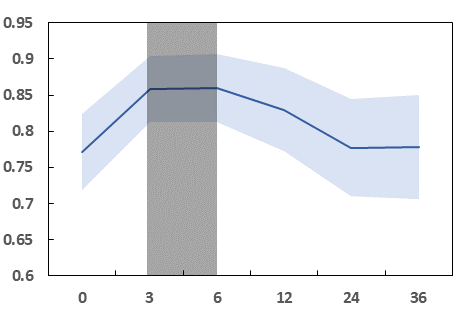
Fig 4 (page 15).
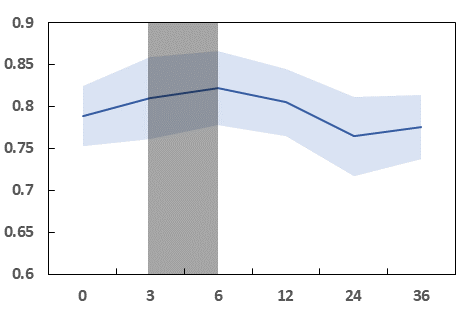
Fig 5 (page 15).
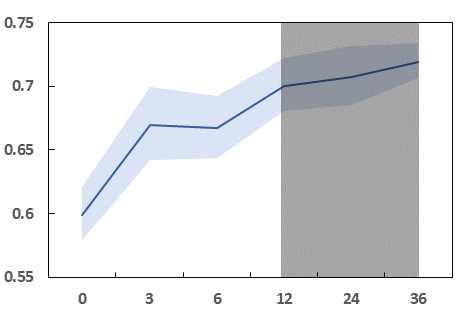
Fig 4: AUROC of the four models for prediction of knee and hip readmission versus
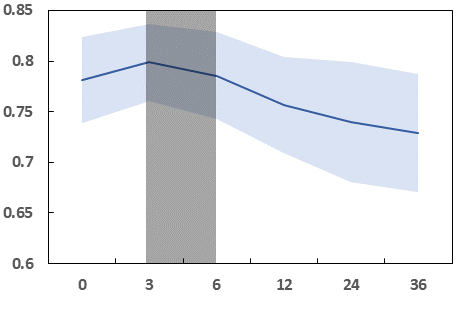
Fig 7 (page 16).
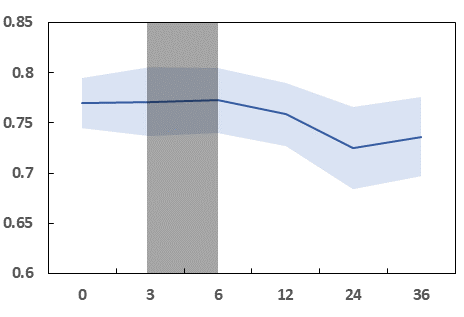
Fig 8 (page 16).
Limitations
- Study data comes from a single healthcare system and EHR provider (Epic), limiting geographical and systemic generalizability.
- The dataset excludes patients without clinical notes, deceased patients, and over 90 years of age, possibly biasing cohort representation.
- No incorporation of expert clinical risk assessment, genomic data, sensor data or other personalized predictors.
- Missing data imputation and categorization of ICD codes may introduce labeling and feature biases.
- Class imbalance remains a challenge with low positive readmission prevalence.
- No external validation or evaluation under domain/distribution shift scenarios.
- Limited transparency on hyperparameter tuning, optimizer choices, and training stability details affects exact reproducibility.
Open questions / follow-ons
- How would these temporal window findings generalize to other surgical or medical readmission scenarios beyond hip and knee arthroplasty?
- What is the impact of incorporating expert clinical risk scores or additional personalized data streams (genomics, sensor data) on temporal data requirements?
- Can model performance and temporal data preferences shift under different EHR systems or broader multi-institutional cohorts?
- How do varying class imbalance handling strategies affect optimal temporal window and prediction robustness?
Why it matters for bot defense
While this study is focused on a clinical prediction problem rather than traditional bot-defense or CAPTCHA systems, the core insight that modality-specific temporal data windows yield different optimal predictive performance has direct conceptual relevance. For example, in security contexts where historical behavior logs versus unstructured user interaction data (like text or clickstreams) might be used, this research highlights the value of tailoring data retention and feature extraction timelines by modality rather than assuming uniformly longer histories improve detection accuracy. Bot-defense engineers should consider whether free-text or interaction logs require fresher data compared to structured event logs, and optimize feature extraction accordingly to improve model efficiency and accuracy. The demonstrated robustness of temporal requirements across model architectures also suggests such design principles may be generalizable beyond healthcare domains. Overall, the paper encourages practitioners to empirically validate time windows rather than defaulting to naive "more history is better" assumptions, which can reduce latency and data storage overhead for real-time threat identification systems.
Cite
@article{arxiv2605_00738,
title={ Temporal Data Requirement for Predicting Unplanned Hospital Readmissions },
author={ Ramin Mohammadi and Vahab vahdat and Sarthak Jain and Amir T. Namin and Ramya Palacholla and Sagar Kamarthi },
journal={arXiv preprint arXiv:2605.00738},
year={ 2026 },
url={https://arxiv.org/abs/2605.00738}
}