UHR-Net: An Uncertainty-Aware Hypergraph Refinement Network for Medical Image Segmentation
Source: arXiv:2604.28095 · Published 2026-04-30 · By Shuokun Cheng, Jinghao Shi, Kun Sun
TL;DR
This paper addresses the challenge of accurate lesion segmentation in medical images, which is complicated by the visual ambiguity of lesions that resemble surrounding tissues and have ill-defined boundaries. Additionally, cues from small lesions tend to be diluted by multi-scale feature extraction in CNNs, leading to under- or over-segmentation. To tackle these problems, the authors propose UHR-Net, an Uncertainty-Aware Hypergraph Refinement Network that integrates two key innovations: (1) an Uncertainty-Oriented Instance Contrastive (UO-IC) pretraining method that uses geometry-aware copy-paste augmentation combined with lesion-like background hard-negative mining to enhance small-lesion discriminability; (2) an Uncertainty-Guided Hypergraph Refinement (UGHR) block that uses an entropy-based uncertainty map derived from a coarse probability segmentation map to modulate node-hyperedge participation in a hypergraph, allowing higher-order interactions to be focused on ambiguous boundary and transition regions.
Experimental evaluation on five public lesion segmentation benchmarks (ISIC-2016, ISIC-2017, GlaS, Kvasir-SEG, Kvasir-Sessile) shows that UHR-Net consistently outperforms strong baseline methods by margins of roughly 1-2% in mIoU and mDSC, demonstrating improved prediction consistency and boundary delineation. Ablations confirm the contribution of each component. Visualization of learned representations shows better separation of small-lesion versus lesion-like background features. Entropy-based uncertainty maps highlight ambiguous regions that guide refinement.
Key findings
- UHR-Net achieves best mIoU scores of 83.5%, 85.0%, and 87.0% on Kvasir-Sessile, Kvasir-SEG, and GlaS datasets respectively, outperforming prior strong baselines by up to 1.9%.
- On ISIC-2016 and ISIC-2017 datasets, UHR-Net attains top mIoU of 87.6% and 81.8%, with corresponding mDSC of 92.9% and 89.2%, showing clear improvements over high-performing methods like ConDSeg and MSCB-UNet.
- Ablation (Table III) shows UO-IC pretraining alone improves mIoU by 1.0% over baseline; enabling base hypergraph refinement adds 0.8%; uncertainty guidance and foreground/background prototype grouping add further improvements, with the full model reaching 85.0% mIoU.
- Optimal hyperedge prototype number M in UGHR is 8 per group, balancing performance gains and complexity (Table IV).
- Logit modulation strength β=1.0 yields peak performance; deviations reduce mIoU slightly (Table V).
- InfoNCE temperature τ=0.10 maximizes contrastive pretraining benefits (Table VI).
- Contrastive loss weight λic=1.0 achieves best balance of segmentation and contrastive objectives during pretraining (Table VII).
- t-SNE visualizations indicate UO-IC pretraining tightens small-lesion feature clusters and separates lesion-like background features more distinctly (Fig 3).
- Entropy-based uncertainty maps derived from coarse segmentation selectively highlight lesion boundaries and ambiguous transition regions, serving as effective guides for refinement (Fig 4).
Threat model
The work does not define an explicit adversary or attack scenario. The implicit ‘adversary’ is the intrinsic ambiguity and similarity between lesion and background pixels in medical images, which causes segmentation errors and unstable predictions. The model assumes a standard supervised learning setting without targeted adversarial manipulation.
Methodology — deep read
Threat model & assumptions: This study assumes an end-to-end supervised segmentation scenario on medical images where lesion boundaries are ambiguous and small lesions are difficult to discriminate. The adversary is implicitly the intrinsic data ambiguity causing unstable predictions. No explicit adversary or attack model is considered.
Data: The authors conduct experiments on five publicly available lesion segmentation datasets: ISIC-2016 and ISIC-2017 for skin lesion segmentation, GlaS for gland segmentation, and Kvasir-SEG and Kvasir-Sessile for gastrointestinal polyp segmentation. The datasets include ground truth masks. They follow standard training/validation/testing splits per prior work. Preprocessing details are not extensively described but presumably include normalization and resizing to network input size.
Architecture / algorithm:
- Backbone network initializes with UO-IC pretraining, which uses geometry-aware lesion copy-paste augmentation to create positive pairs (original lesion and downscaled replica) and mines lesion-like background regions as hard negatives based on prediction confidence maps.
- Contrastive learning is performed with an InfoNCE loss on pooled instance features extracted by masked average pooling from the backbone feature maps.
- During end-to-end training, a lightweight guidance head produces a coarse segmentation probability map.
- The UGHR block refines features through uncertainty-guided hypergraph operations:
- Uncertainty map U is computed as pixel-wise normalized entropy from the coarse probability map.
- Nodes represent pixels; two groups of hyperedge prototypes are dynamically generated for foreground and background conditioned on the coarse priors.
- Node-hyperedge participation logits are computed by scaled dot product between node features and prototypes, modulated by uncertainty values to emphasize ambiguous nodes.
- Participation weights are normalized and used to perform message passing between nodes and hyperedges, updating node features.
- A parallel dilated convolutional branch locally enhances features.
- Refined features combine outputs from both branches.
- Training regime:
- UO-IC pretraining uses batch size 16, Adam optimizer with lr=1e-4, contrastive temperature τ=0.10, loss weight λic=1.0.
- End-to-end training uses the same batch size and optimizer settings, with auxiliary loss weight λaux=0.1 to supervise the coarse probability map.
- The hypergraph prototype number M=8 per group, modulation strength β=1.0.
- The full training involves jointly optimizing segmentation (Dice + BCE loss), contrastive loss during pretraining, and auxiliary coarse segmentation supervision during full training.
- Evaluation protocol:
- Metrics include mean Intersection over Union (mIoU), mean Dice similarity coefficient (mDSC), recall and precision.
- Comparisons are made against multiple recent strong segmentation baselines such as U-Net++, ESPNet, ConDSeg, MSCB-UNet, BGDiffSeg etc.
- Ablation studies analyze the effect of UO-IC pretraining, hypergraph components (base HR, uncertainty guidance, foreground/background prototype split).
- Sensitivity to hyperparameters like M, β, τ, and λic is explored.
- Qualitative visualizations show uncertainty maps and t-SNE of learned feature embeddings.
- Reproducibility: The authors provide code at https://github.com/CUGfreshman/UHR-Net. All datasets are public benchmarks. Detailed training hyperparameters and network architectures are described in the paper, supporting replication though pretrained weights are not explicitly mentioned.
Example end-to-end flow: Given an input image, during pretraining, geometry-aware copy-paste augmentation creates an image with a scaled lesion pasted into a lesion-free region, generating positive and negative feature pairs extracted via masked pooling. The InfoNCE contrastive loss trains the backbone to better discriminate lesions from lesion-like backgrounds. During full training, the backbone extracts multi-scale features, while a lightweight guidance head predicts a coarse probability map. The entropy-based uncertainty map derived from this coarse map modulates node-hyperedge interactions in multiple UGHR blocks. Foreground and background hyperedge prototypes are conditioned on the coarse map through cross-attention and adapted for each sample. Participation weights are modulated by uncertainty to focus refinement on ambiguous regions, and hypergraph message passing propagates contextual information to improve boundary prediction and stability. The refined multi-scale features are decoded to produce the final segmentation mask, trained using Dice+BCE loss with auxiliary supervision on the coarse map.
Technical innovations
- Uncertainty-Oriented Instance Contrastive pretraining combines geometry-aware lesion copy-paste augmentation with lesion-like background hard-negative mining to enhance instance-level discriminative features for small, ambiguous lesions.
- Uncertainty-Guided Hypergraph Refinement block uses an entropy-based uncertainty map derived from coarse segmentation probabilities to modulate node-hyperedge participation logits, emphasizing ambiguous regions for targeted feature refinement.
- Dynamic generation of foreground- and background-conditioned hyperedge prototype groups with bidirectional cross-attention enables decoupled higher-order message passing to reduce foreground-background interference.
- Integration of contrastive pretraining with structured hypergraph refinement in an end-to-end segmentation network tailored for medical lesion segmentation, balancing instance-level discrimination and pixel-level contextual refinement.
Datasets
- ISIC-2016 — ~900 images — public skin lesion segmentation dataset
- ISIC-2017 — ~2000 images — public skin lesion segmentation dataset
- GlaS — ~165 images — public gland segmentation dataset from histology
- Kvasir-SEG — 1000 polyp images — public gastrointestinal endoscopy dataset
- Kvasir-Sessile — subset of Kvasir-SEG — public gastrointestinal dataset
Baselines vs proposed
- Kvasir-Sessile: ConDSeg mIoU = 81.2% vs UHR-Net = 83.5%
- Kvasir-SEG: ESPNet mIoU = 83.6% vs UHR-Net = 85.0%
- GlaS: ConDSeg mIoU = 85.1% vs UHR-Net = 87.0%
- ISIC-2016: ConDSeg mIoU = 86.8% vs UHR-Net = 87.6%
- ISIC-2017: MSCB-UNet mIoU = 81.2% vs UHR-Net = 81.8%
- Ablation (Table III): Baseline mIoU 81.2%, +UO-IC 82.2%, +Base HR 83.0%, +FG/BG HG 83.5%, +Unc. Guid. 83.9%, full model 85.0%
Figures from the paper
Figures are reproduced from the source paper for academic discussion. Original copyright: the paper authors. See arXiv:2604.28095.

Fig 1: Overall framework of the proposed UHR-Net. The upper left part is the UO-IC pretraining stage. The lower left part is the end-to-end training

Fig 2 (page 3).

Fig 3 (page 3).

Fig 4 (page 3).

Fig 5 (page 3).
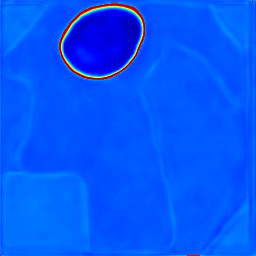

Fig 6 (page 3).
Fig 2: Qualitative comparisons on ISIC-2016 and Kvasir-SEG. Columns show Image, U-Net++, ESPNet, CMUNeXt, ConDSeg, Ours, and Ground Truth
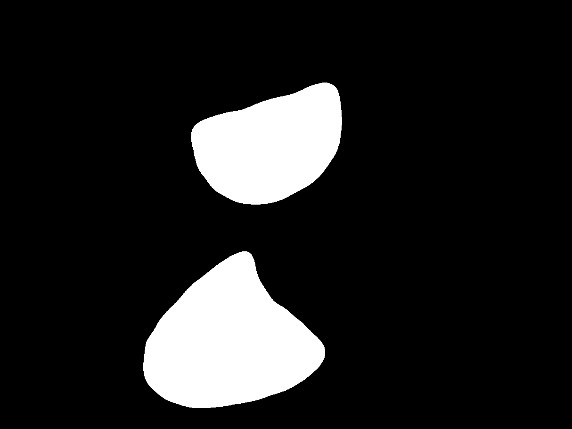

Fig 3: presents a t-SNE visualization of three types of
Limitations
- The method focuses on five public datasets mostly of skin lesions and gastrointestinal polyps; generalization to other lesion types or imaging modalities is not studied.
- No explicit adversarial robustness or attack evaluation is performed to test resilience against malicious perturbations.
- Computational complexity and inference speed overhead of UGHR blocks compared to baselines are not thoroughly quantified.
- The approach relies on accurate coarse segmentation outputs to generate uncertainty maps; failure modes when coarse maps are very poor are not analyzed.
- Uncertainty guidance is derived from entropy of coarse predictions only; other uncertainty modeling paradigms (e.g., Bayesian) are not explored.
- While instance-level contrastive pretraining improves small lesion discrimination, it requires manual lesion instance extraction and geometric transformations that may not transfer easily.
Open questions / follow-ons
- How would the uncertainty-guided hypergraph refinement perform under domain shift or unseen lesion types, and can it adapt dynamically?
- Can the UO-IC pretraining strategy be generalized to other medical segmentation tasks beyond lesions, or to 3D volumetric data?
- Would integrating probabilistic uncertainty estimation methods (e.g., MC-dropout, ensembles) improve or complement the entropy-based uncertainty maps?
- What is the tradeoff in inference speed and memory usage introduced by multi-scale UGHR blocks in large-scale clinical deployment?
Why it matters for bot defense
For bot-defense engineers working on CAPTCHA or automated detection systems, this paper illustrates how uncertainty quantification—specifically entropy-based uncertainty maps—can be directly integrated into structured refinement modules (here via hypergraphs) to focus processing resources on ambiguous or hard-to-classify regions. This principle may translate to improving bot-detection models that encounter borderline or adversarial examples by emphasizing uncertain regions for further scrutiny. Moreover, the UO-IC contrastive pretraining shows the benefit of explicitly mining hard negative examples and nearby ambiguous instances to strengthen discrimination of subtle cues, which analogously could be applied to distinguishing sophisticated bots from humans that mimic legitimate behavior.
The hypergraph formulation moving beyond pairwise relationships to higher-order context aggregation is also relevant for representing complex dependencies in behavioral or interaction features. However, practical application would require adaptation to temporal or networked data common in bot-detection. The paper’s methodology highlights the value of combining uncertainty estimation, instance-level representation learning, and higher-order relational reasoning to enhance robustness and precision in challenging classification scenarios involving ambiguous samples.
Cite
@article{arxiv2604_28095,
title={ UHR-Net: An Uncertainty-Aware Hypergraph Refinement Network for Medical Image Segmentation },
author={ Shuokun Cheng and Jinghao Shi and Kun Sun },
journal={arXiv preprint arXiv:2604.28095},
year={ 2026 },
url={https://arxiv.org/abs/2604.28095}
}